Chapter 4 Reproducibility
by Jade Benjamin-Chung
Our lab adopts the following practices to maximize the reproducibility of our work.
- Design studies with appropriate methodology and adherence to best practices in epidemiology and biostatistics
- Register study protocols
- Write and register pre-analysis plans
- Create reproducible workflows
- Process and analyze data with internal replication and masking
- Use reporting checklists with manuscripts
- Publish preprints
- Publish data (when possible) and replication scripts
4.1 What is the reproducibility crisis?
In the past decade, an increasing number of studies have found that published study findings could not be reproduced. Researchers found that it was not possible to reproduce estimates from published studies: 1) with the same data and same or similar code and 2) with newly collected data using the same (or similar) study design. These “failures” of reproducibility were frequent enough and broad enough in scope, occurring across a range of disciplines (epidemiology, psychology, economics, and others) to be deeply troubling. Program and policy decisions based on erroneous research findings could lead to wasted resources, and at worst, could harm intended beneficiaries. This crisis has motivated new practices in reproducibility, transparency, and openness. Our lab is committed to adopting these best practices, and much of the remainder of the lab manual focuses on how to do so.
The scientific process involves a series of steps from defining a hypothesis through publishing results, and threats to reproducibility can arise at each stage. The figure below summarizes common threats that can occur at each step in the scientific process, including bias and low statistical power in study design, measurement error during data collection, and confirmation bias, p-hacking, and errors during analysis.
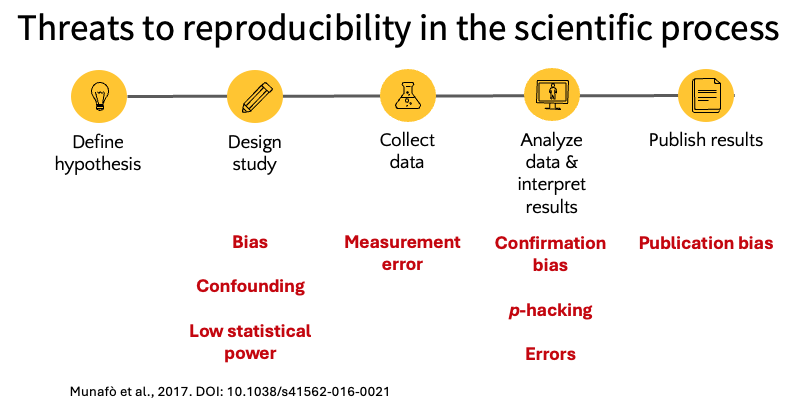
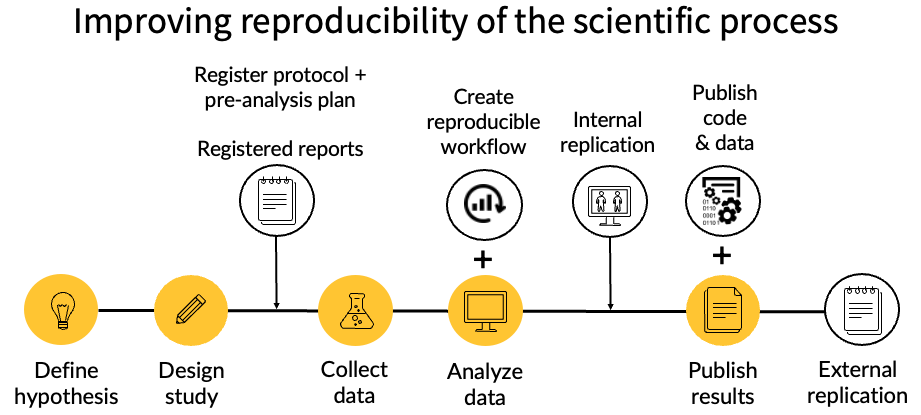
Our lab attempts to minimize these threats through a set of practices at each stage of the research process. The figure below illustrates our approach: we register protocols and pre-analysis plans before data collection, create reproducible computational workflows and conduct internal replication during analysis, and publish code and data alongside our results to enable external replication. The rest of this chapter describes each of these practices in detail.
Recommended readings on the “reproducibility crisis”:
Nuzzo R. How scientists fool themselves – and how they can stop. 2015. https://www.nature.com/articles/526182a
Stoddart C. Is there a reproducibility crisis in science? 2016. https://www.nature.com/articles/d41586-019-00067-3
Munafo MR, et al. A manifesto for reproducible science. Nature Human Behavior 2017 http://dx.doi.org/10.1038/s41562-016-0021
4.2 Study design
Appropriate study design is beyond the scope of this lab manual and is something trainees develop through their coursework and mentoring.
4.3 Register study protocols
We register all randomized trials on clinicaltrials.gov, and in some cases register observational studies as well.
4.4 Write pre-analysis plans
We write pre-analysis plans for most original research projects that are not exploratory in nature, although in some cases, we write pre-analysis plans for exploratory studies as well. The format and content of pre-analysis plans can vary from project to project. Generally, these include:
- Brief background on the study (a condensed version of the introduction section of the paper)
- Hypotheses / objectives
- Study design
- Description of data
- Definition of outcomes
- Definition of interventions / exposures
- Definition of covariates
- Statistical power calculation
- Statistical analysis:
- Type of model
- Covariate selection / screening
- Standard error estimation method
- Missing data analysis
- Assessment of effect modification / subgroup analyses
- Sensitivity analyses
- Negative control analyses
See the following pre-analysis plans for examples from our lab:
4.5 Register pre-analysis plans
We use Open Science Framework to register pre-analysis plans. Learn more here. Briefly, what this entails is creating a time-stamped, publicly accessible record of your hypotheses, study design, and analysis plan before you analyze the data. Registration does not prevent you from conducting additional analyses — it simply makes transparent which analyses were planned in advance and which were exploratory. To register, create a project on OSF, upload your pre-analysis plan document, and select “Register” to create a frozen, time-stamped version. Once registered, the plan cannot be edited or deleted, though you can add addenda if your analysis plan changes after registration. You can also include a link to the registration at the time of submitting your manuscript. This shows peer reviewers and editors (and later, readers) that you pre-specified your analysis.
People often ask, well what if I need to change my plan after pre-specifying it? No problem! We commonly include a list of deviations from the pre-analysis plan in the supplement of our manuscript. In our experience, this has only strengthened reviewer and editor perceptions of the reproducibility and rigor of our work because it shows that we are transparent about our process and that we are not hiding any changes to the analysis plan. We have even included a list of deviations resulting from the peer review process as its own section in the supplement to make those changes transparent as well.
4.6 Create reproducible workflows
Reproducible workflows allow a user to reproduce study estimates and ideally figures and tables with a “single click”. In practice, this typically means running a single bash script that sources all replication scripts in a repository. These replication scripts complete data processing, data analysis, and figure/table generation. The following chapters provide detailed guidance on this topic:
- Getting started: Overview of the tools used in our lab
- Code repositories: Project structure and configuration
- Coding practices: Script organization, documentation, and code review
- Coding style: Formatting conventions
- Data validation and unit testing: Assertions, sanity checks, and unit testing
- Working with big data: Handling large datasets
- Tidy evaluation and programming with dplyr: Programming with tidyverse functions
- GitHub: Version control and collaboration
- Unix commands: Running scripts from the command line
- Reproducible environments: Package and R version management
- Code publication: Preparing code for public release
4.7 Process and analyze data with internal replication and masking
See my video on this topic: https://www.youtube.com/watch?v=WoYkY9MkbRE
4.8 Use reporting checklists with manuscripts
Using reporting checklists helps ensure that peer-reviewed articles contain the information needed for readers to assess the validity of your work and/or attempt to reproduce it. A collection of reporting checklists is available here: https://www.equator-network.org/about-us/what-is-a-reporting-guideline/)
4.9 Publish preprints
A preprint is a scientific manuscript that has not been peer reviewed. Preprint servers create digital object identifiers (DOIs) and can be cited in other articles and in grant applications. Because the peer review process can take many months, publishing preprints prior to or during peer review enables other scientists to immediately learn from and build on your work. Importantly, NIH allows applicants to include preprint citations in their biosketches. In most cases, we publish preprints on medRxiv.
4.10 Publish data (when possible) and replication scripts
Publishing data and replication scripts allows other scientists to reproduce your work and to build upon it. We typically publish data on Open Science Framework, share links to Github repositories, and archive code on Zenodo.